
So nat’ralists observe, a flea
Hath smaller fleas that on him prey;
And these have smaller fleas to bite ‘em.
And so proceeds Ad infinitum.
Jonathan Swift, 1733
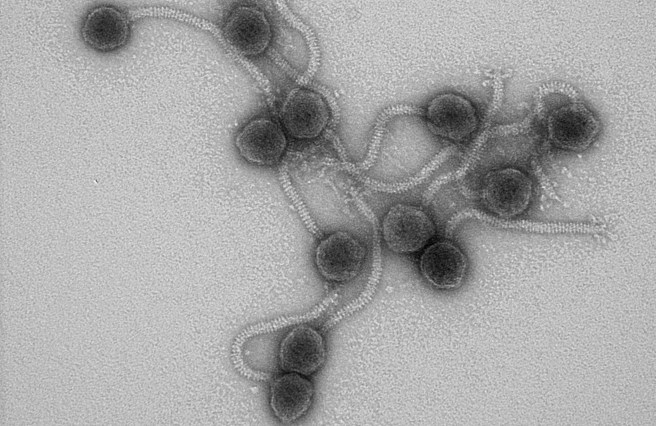
When the Anglo-Irish satirist wrote these words nearly two centuries ago, he could not have known just how far down the tree of life his observations would hold true. These predator-prey relationships exist beyond the plains of Africa or the jungles of Borneo. They extend to the realm of microscopic organisms and to the world of bacteria and the teeny tiny, itsy bitsy viruses that prey on them. These viruses are called bacteriophages, or phages for short.
Like human and other animal viruses, phages rely completely on their host for reproduction. They enter a bacterial cell and hijack the cellular machinery to make new phages until the cell is literally bursting with viral cargo. A torrent of phages is unleashed that go on to infect more bacteria and continue the cycle.
But bacteria are not helpless victims in this story. They have a large arsenal of anti-phage weapons to keep phages out and prevent them from taking over. Perhaps the coolest of these weapons is the CRISPR-Cas system. First discovered in 2007, the CRISPR-Cas system functions as the bacteria’s immune system. It is both a memory keeper and a hitman. Every time a bacteria survives a phage infection (which doesn’t happen often), the CRISPR-Cas complex takes a small piece of phage DNA and adds it to the bacteria’s own DNA, gradually building a database of unique DNA fingerprints from every phage that has ever tried to kill it. In other words, bacteria with CRISPR-Cas systems are able to “learn” from their previous phage encounters and acquire immunological memory based on those experiences—a trait that was previously thought to be unique to animals.
“It’s the first example of a single cell, simple [bacteria] having an adaptive immune system,” says Dr. Joseph Bondy-Denomy, a faculty fellow at the University of California, San Francisco. “The adaptability of CRISPR is very, very rapid. I think that’s why it’s so exciting.”
Each CRISPR-Cas complex comprises different proteins and a single piece of RNA that matches a unique phage DNA fingerprint. The complexes act like patrol cars driving around, each searching for a different suspect based on a different “WANTED” poster. The next time one of these phages tries to infect the bacteria, the CRISPR-Cas complex recognizes it based on a DNA match and chops up the incoming phage’s DNA, effectively destroying it before it has a chance to harm its host.
Three years ago, Bondy-Denomy, then a PhD student at the University of Toronto, discovered that some phages carried genes that could disable their bacterial host’s CRISPR-Cas system. He was studying phages that infect Pseudomonas aeruginosa, a ubiquitous soil bacteria that can cause persistent lung infections in patients with cystic fibrosis. Although the genome of P. aeruginosa contained all the genes required for two different CRISPR-Cas systems, researchers didn’t think the systems worked because they had never seen it prevent phage infections. From the 80 new phages he identified for an earlier part of his project, Bondy-Denomy stumbled upon a few phages that were not able to kill their bacterial hosts. He later showed that unlike the majority of phages in his collection, these select phages were being targeted by the CRISPR-Cas complex and blocked from replicating in the bacterial cell.
“That told us right away that the CRISPR system is actually functional [in P. aeruginosa],” says Bondy-Denomy. “It seems that it wasn’t the [CRISPR-Cas system] of the bacteria that was faulty. It was the phages could actively stop the CRISPR system from working.” Eventually, he identified the phage genes and proteins responsible for disarming P. aeruginosa’s CRISPR-Cas system. He called these genes anti-CRISPRs.
Anti-CRISPRs represent the latest escalation in the ancient arms race between bacteria and phages. In a new paper published this week in Nature, Bondy-Denomy and colleagues at the University of Toronto and Montana State University have shown how these anti-CRISPRs inhibit the CRISPR-Cas complex.
“The initial anti-CRISPR discovery was perplexing because it wasn’t just a single gene—there were about nine of them that could each turn off two different types of CRISPR [systems on its own],” he says. On top of that, these nine genes looked nothing like each other. “The main question was, how do they work?” says Bondy-Denomy. “Are they somehow all doing the same thing to turn off the CRISPR system? Or are they all performing the same function, but in a different way?”
In their paper, the researchers focused on three different anti-CRISPR proteins and found that they inhibited the CRISPR-Cas system in three different ways. Two of the anti-CRISPRs prevented the CRISPR-Cas complex from grabbing phage DNA by attaching themselves to different parts of the complex. The third anti-CRISPR blocked the protein that functions as a pair of molecular scissors to cut up phage DNA.
For Bondy-Denomy, these results are both surprising and exciting—surprising because these are three completely different genes that do completely different things at the molecular level to produce the same outcome; and exciting because this is the first time that proteins that directly engage with the CRISPR-Cas complex have been identified. Human cells also use protein-RNA complexes similar to CRISPR-Cas to survey the environment for foreign invaders. What’s more, researchers have shown that by interacting with different protein partners, these complexes can be made to have alternative functions like turning on or off certain human genes. By showing that the anti-CRISPR proteins directly interact with the CRISPR-Cas complex, Bondy-Denomy and his colleagues have uncovered the tantalizing new prospect of finding more interactors that could unlock new functions for CRISPR inside the cell.
In the constant battle between bacteria and phages, one might expect bacteria to counter a phage’s anti-CRISPR with an anti-anti-CRISPR. Should we expect to learn about that soon? “It’s kind of a mystery what that is right now,” says Bondy-Denomy. “It’s a very hard question to answer but it’s fun to think about.”
References:
Bondy-Denomy J, Pawluk A, Maxwell KL, & Davidson AR (2013). Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature, 493 (7432), 429-32 PMID: 23242138
Bondy-Denomy, J., Garcia, B., Strum, S., Du, M., Rollins, M., Hidalgo-Reyes, Y., Wiedenheft, B., Maxwell, K., & Davidson, A. (2015). Multiple mechanisms for CRISPR–Cas inhibition by anti-CRISPR proteins Nature DOI: 10.1038/nature15254

Great summary and explanation – very well written. So did you get in contact with Dr Bondy-Denomy to discuss his research for this post?
LikeLiked by 1 person
Thanks Lucy! Glad you like it 🙂 Yes, I did speak with Dr. Bondy-Denomy to talk about his work. I linked to his lab website in the post so you can find his contact info there. He’s very friendly!
LikeLiked by 1 person